News

INDICATE Project receives a grant from the Wilhelm Sander Foundation
Funding for INDICATE
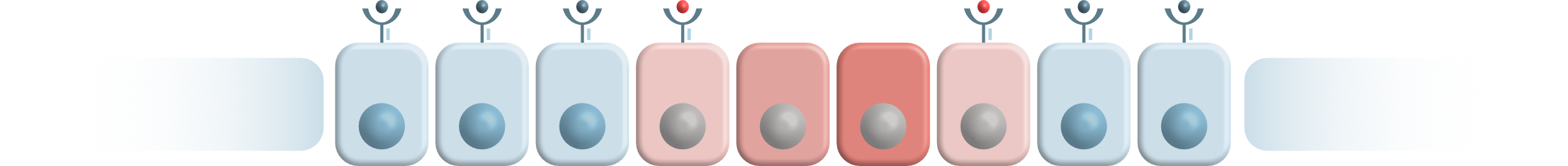
We hear a lot about environmental factors influencing our cancer risk. But what about our immune system? The INDICATE project offers a chance to understand the intricate associations between host’s immune constitution and cancer risk by focusing on a population with increased cancer risk due to genetic predisposition. And now, with the generous support of the Wilhelm Sander Foundation, INDICATE gets a great momentum!
INDICATE is an international scientific network aiming at analyzing the role of HLA type in Lynch syndrome-associated cancer risk. By collecting data from across the world we want to elucidate whether HLA type is a risk modifier in Lynch syndrome. A better understanding of risk modifiers will help us to precisely estimate individual cancer risk and enable risk-adapted cancer prevention strategies.
We are very grateful to the Wilhelm Sander Foundation directed by Bernard Knappe for generously funding this project at the Department of Applied Tumor Biology, Heidelberg University Hospital for a duration of two years.
INDICATE initiative highlighted in Ärzteblatt
We are very grateful to Ärzteblatt for publishing our study announcement in their current issue.
This is a great chance for INDICATE to expand our outreach and involve as much researchers, scientists, clinicians and patients interested in our study as possible, to collaborate on immune factors determining tumor risk in Lynch syndrome.
INDICATE Consortium publishes its first collaborative paper on the role of HLA type in determining cancer risk and announces the launch of the INDICATE initiative
© 2022 The Authors. International Journal of Cancer published by John Wiley & Sons Ltd on behalf of Union for International Cancer Control
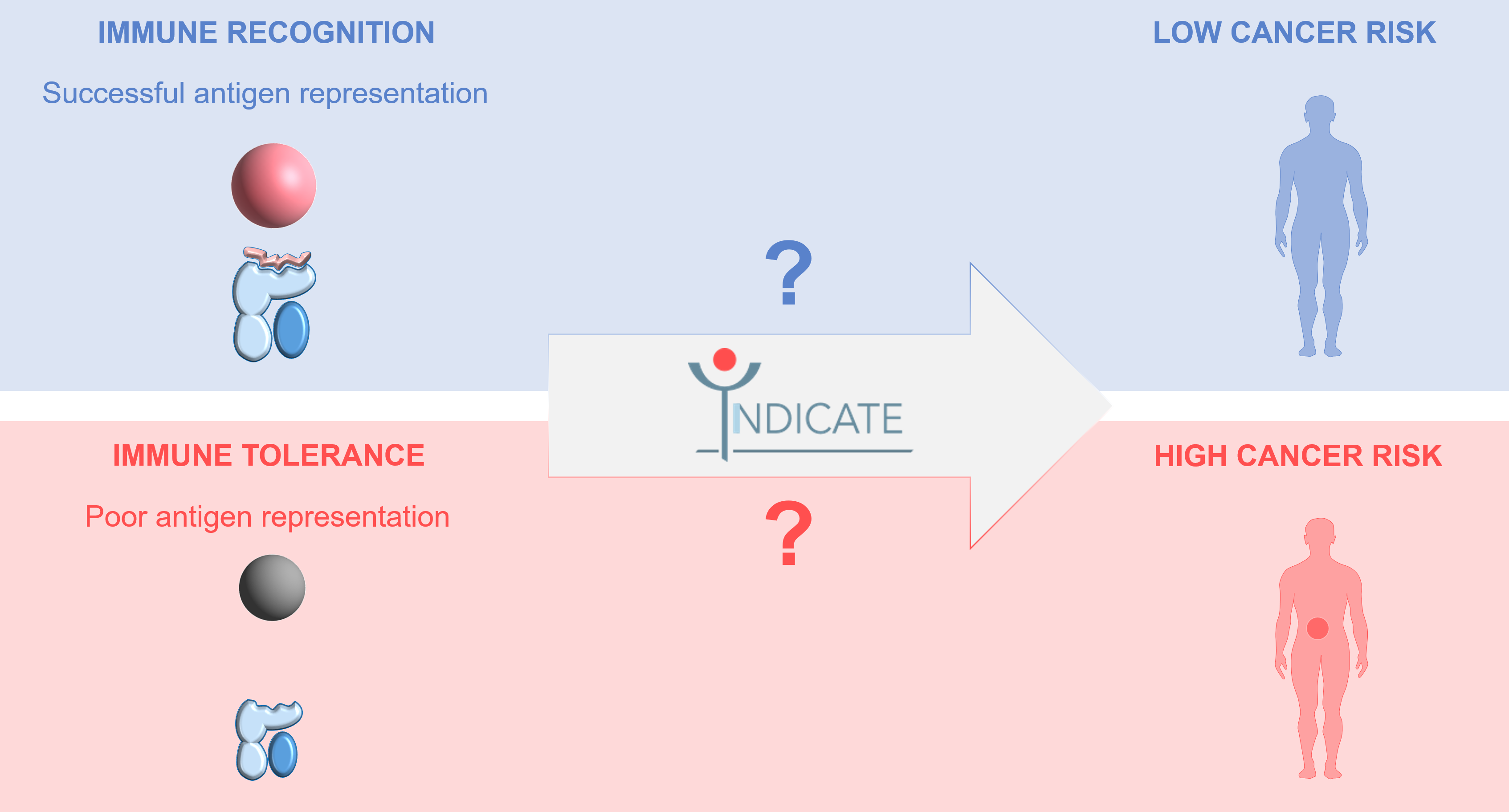
The associations of certain HLA types with disease susceptibility have been previously shown for virus infections, however in cancer this aspect remains poorly understood. Lynch syndrome with its genetically defined population and strong involvement of the immune system in the carcinogenic process is an ideal model to address this question for the first time in a systematic manner.
Our previous research pointed at a possible influence of the HLA type on the potential of the immune system to eliminate cancer cells. Together with our colleagues from DMQ (HITS) and our INDICATE partners from Germany, the United Kingdom, Finland, the Netherlands, Norway, and Hungary we now have published an open access article in the International Journal of Cancer, describing the fascinating interplay between the HLA system and human disease susceptibility, announcing the launch of INDICATE and inviting researchers worldwide to collaborate in this project.
Our new paper describing a methodological approach for detecting certain HLA alleles in archival tissue samples is published in HLA
© 2022 The Authors. HLA published by John Wiley & Sons Ltd on behalf of Union for International Cancer Control
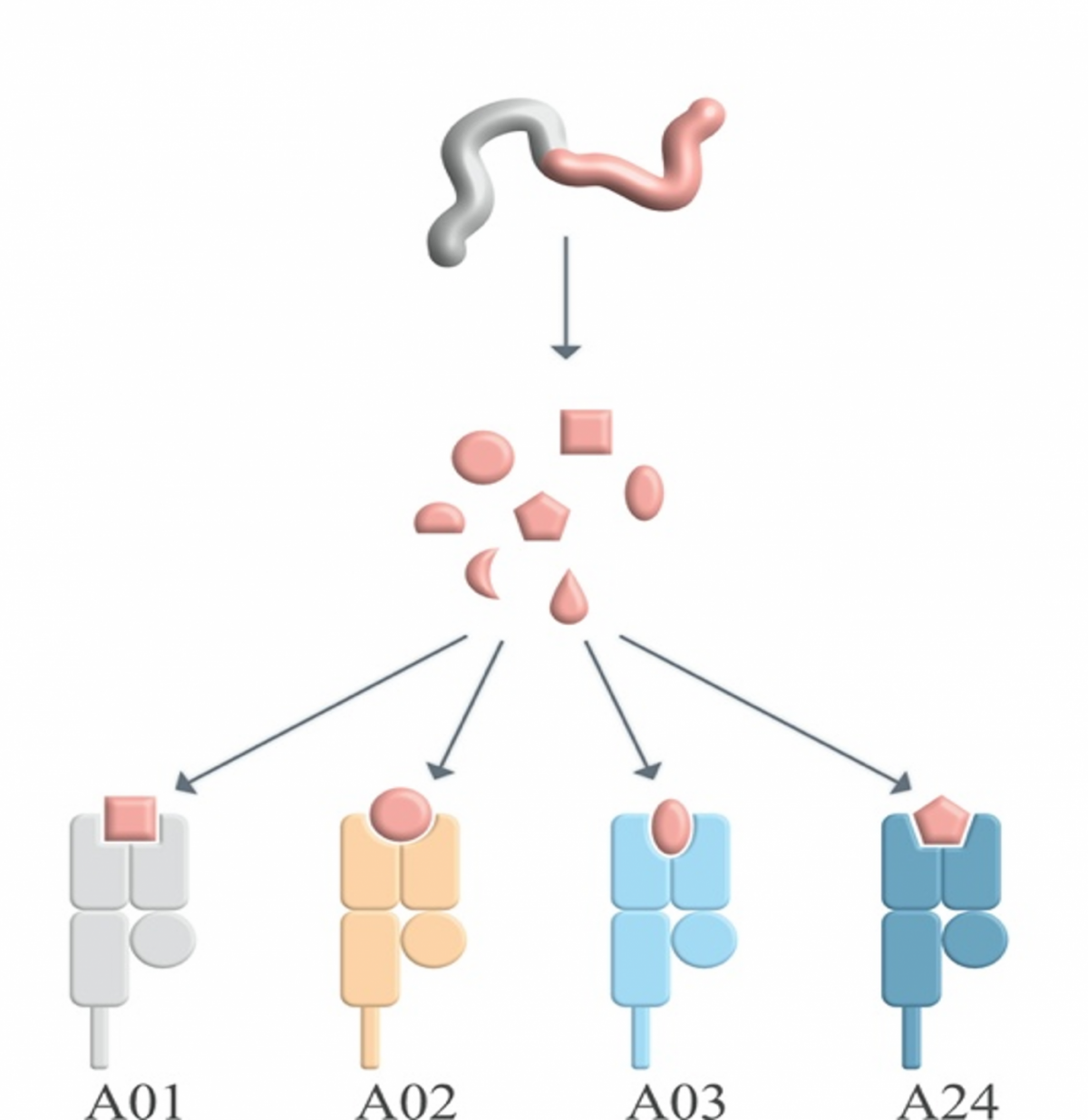
Particularly in cancer research, a major type of study material is archival formalin-fixed paraffin-embedded tissue. However, the DNA in this type of tissue is often fragmented and not suitable for certain analyses, such as determining HLA alleles. To enable the analysis of certain HLA alleles in this type of material and make it available for research, we developed a refined approach for characterizing presence or absence of HLA-A*02, the most common HLA-A allele in the Caucasian population, in archival samples, which is now published in HLA. By sharing this methods with our peers, we want to promote studying HLA type-dependency during the pathogenesis of a wide range of diseases and make archival and historic tissue samples accessible for identifying HLA-A*02 alleles.



